AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Hypnogram unit of measurement2/8/2024 sum () if scaled : # Handle division by zero error denom = x. name = "Channel" def _coincidence ( x, y ): """Calculate the (scaled) coincidence.""" coincidence = ( x * y ). _ch_names ) < 2 : raise ValueError ( "At least 2 channels are required to calculate coincidence." ) mask = self. concat ( output, ignore_index = True ) return output def get_coincidence_matrix ( self, scaled = True ): """get_coincidence_matrix""" if len ( self. append ( df_chan ) if as_dataframe : output = pd. iloc df_chan = i # Append to master dataframe output. iloc ) # Append channel name df_chan = ev_chan. melt ( id_vars = "Time", var_name = "Event", value_name = "Amplitude" ) # Append stage if "Stage" in masked_events : df_chan = df_chan. T ) df_chan = time # Convert to long-format df_chan = df_chan. append ( amps ) continue # Convert to long-format dataframe df_chan = pd. Skipping channel." ) continue # Get data at indices and time vector amps = data if not as_dataframe : # Output is a list (n_channels) of numpy arrays (n_events, n_times) output. error ( "Time before and/or time after exceed data bounds, please " "lower the temporal window around center. to_numpy () # Get centered indices idx, idx_valid = get_centered_indices ( data, peaks, bef, aft ) # If no good epochs are returned raise a warning if len ( idx_valid ) = 0 : logger. unique (): # Copy is required to merge with the stage later on ev_chan = masked_events = i ]. _check_mask ( mask ) masked_events = self. _sf, l_freq = filt, h_freq = filt, method = "fir", verbose = False ) else : data = self. arange ( - bef, aft + 1, dtype = "int" ) / self. Should we add resample=100 (Hz) or step_size=0.01? time = np. info ( "Peak-to-peak amplitude of %s = %.4f uV" % ( ch_names, all_ptp )) if check_amp and not ( 0.1 = 0 assert time_after >= 0 bef = int ( self. info ( "Trimmed standard deviation of %s = %.4f uV" % ( ch_names, all_trimstd )) logger. zeros ( n_chan, dtype = bool ) for i in range ( n_chan ): logger. ptp ( data, axis =- 1 ) all_trimstd = trimbothstd ( data, cut = 0.05 ) bad_chan = np. info ( "Data duration = %.2f seconds", n_samples / sf ) all_ptp = np. info ( "Sampling frequency = %.2f Hz", sf ) logger. info ( "Number of samples in data = %i ", n_samples ) logger. any (), ( "None of the stages specified " "in `include` are present in " "hypno." ) # 4) Check data amplitude logger. kind, "hypno and include must have same dtype" assert np. size >= 1, "`include` must have at least one element." assert hypno. size ) assert include is not None, "include cannot be None if hypno is given" include = np. info ( "Number of unique values in hypno = %i ", unique_hypno. size = n_samples, "Hypno must have same size as data." unique_hypno = np. ndim = 1, "Hypno must be one dimensional." assert hypno. 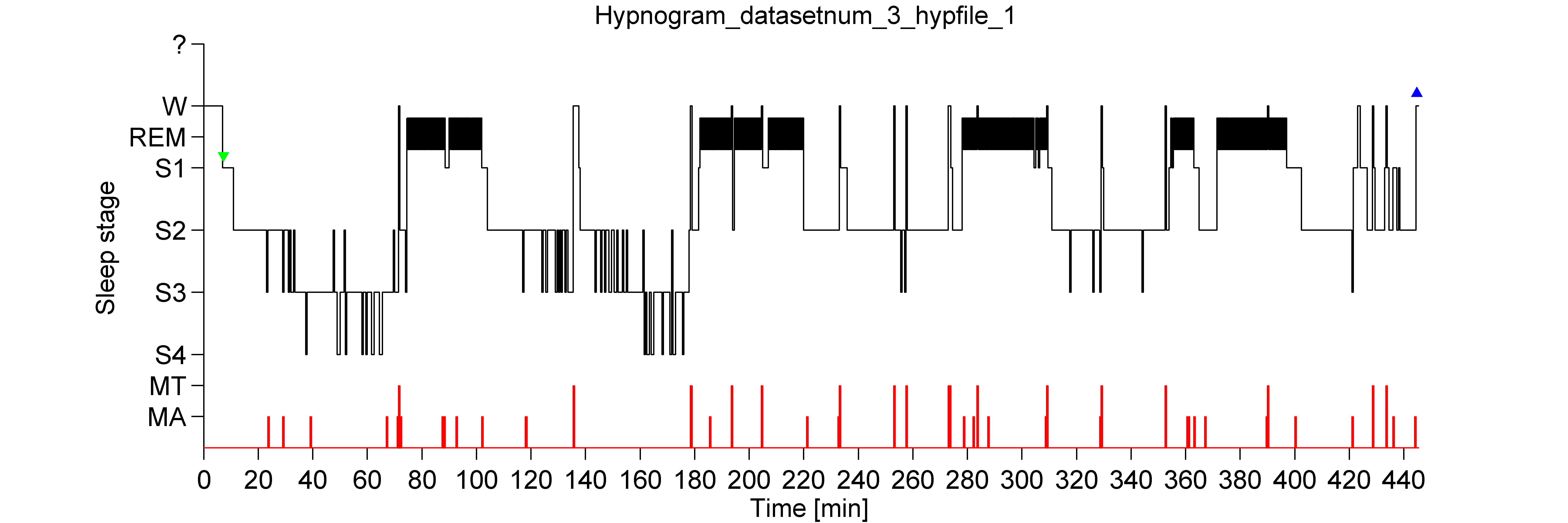
asarray ( hypno, dtype = int ) assert hypno. shape # 2) Check channel names if ch_names is None : ch_names = else : assert len ( ch_names ) = n_chan # 3) Check hypnogram if hypno is not None : hypno = np. ndim = 1 : # Force to 2D array: (n_chan, n_samples) data = data n_chan, n_samples = data. ndim in, "data must be 1D (times) or 2D (chan, times)." if data. ndarray ): # Deal with sf = array(100.) -> 100 sf = float ( sf ) assert isinstance ( sf, ( int, float )), "sf must be int or float." data = np. get_data ( units = dict ( eeg = "uV", emg = "uV", eog = "uV", ecg = "uV" )) else : assert sf is not None, "sf must be specified if not using MNE Raw." if isinstance ( sf, np. ch_names # Extract channel names data = data. info # Extract sampling frequency ch_names = data. getLogger ( "yasa" ) _all_ = # DATA PREPROCESSING # def _check_data_hypno ( data, sf = None, ch_names = None, hypno = None, include = None, check_amp = True ): """Helper functions for preprocessing of data and hypnogram.""" # 1) Extract data as a 2D NumPy array if isinstance ( data, mne. others import ( moving_transform, trimbothstd, get_centered_indices, sliding_window, _merge_close, _zerocrossings, ) logger = logging. io import set_log_level, is_tensorpac_installed, is_pyriemann_installed from. Author: Raphael Vallat (- GitHub: - License: BSD 3-Clause License """ import mne import logging import numpy as np import pandas as pd from scipy import signal from mne.filter import filter_data from collections import OrderedDict from scipy.interpolate import interp1d from scipy.fftpack import next_fast_len from sklearn.ensemble import IsolationForest from. """ YASA (Yet Another Spindle Algorithm): fast and robust detection of spindles, slow-waves, and rapid eye movements from sleep EEG recordings.
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
